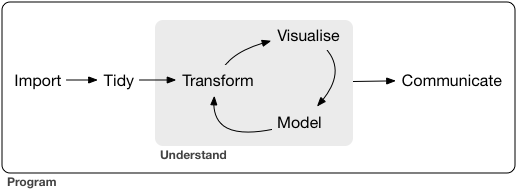
DSCI 100 - Introduction to Data Science¶
Lecture 3 - Wrangling to get tidy data¶
Credit to Jenny Brian's slides and Garrett Grolemund's tidying example
Housekeeping¶
- Reminder about worksheets/tutorials:
- do not ever rename, move, or delete your worksheet
- do not use "Save As..." when you're saving your work
- use "Save Notebook" or ⌘-s / Ctrl-s instead!
- at a minimum, if you move/delete/rename worksheets, our autograder won't be able to find them and will give you 0
- You can download your worksheets and tutorials to your computer if you think something has gone terribly wrong.
Data Wrangling!¶

The cartoon illustrations in this slide deck are all created by Allison Horst
- In the real world, when you get data, it's usually very messy
- inconsistent format (commas, tabs, semicolons, missing data, extra empty lines)
- split into multiple files (e.g. yearly recorded data over many years)
- corrupted files, custom formats
- when you read it successfully into R, it will often still be very messy
- you need to make your data "tidy"
What is Tidy Data?¶

Illustrations from the Openscapes blog Tidy Data for reproducibility, efficiency, and collaboration by Julia Lowndes and Allison Horst"


Examples of Tidy(?) Data¶
...here is the same data represented in a few different ways. Let's vote on which are tidy!
Tuberculosis data¶
This data is tidy. True or false?
| country | year | rate |
|---|---|---|
| Afghanistan | 1999 | 745/19987071 |
| Afghanistan | 2000 | 2666/20595360 |
| Brazil | 1999 | 37737/172006362 |
| Brazil | 2000 | 80488/174504898 |
| China | 1999 | 212258/1272915272 |
| China | 2000 | 213766/1280428583 |
Tuberculosis data¶
This data is tidy. True or false?
| country | cases (year=1999) | cases (year=2000) | population (year=1999) | population (year=2000) |
|---|---|---|---|---|
| Afghanistan | 745 | 2666 | 19987071 | 20595360 |
| Brazil | 37737 | 80488 | 172006362 | 174504898 |
| China | 212258 | 213766 | 1272915272 | 1280428583 |
Tuberculosis data¶
This data is tidy. True or false?
| country | 1999 | 2000 |
|---|---|---|
| Afghanistan | 745 | 2666 |
| Brazil | 37737 | 80488 |
| China | 212258 | 213766 |
Tuberculosis data¶
This data is tidy. True or false?
| country | year | key | value |
|---|---|---|---|
| Afghanistan | 1999 | cases | 745 |
| Afghanistan | 1999 | population | 19987071 |
| Afghanistan | 2000 | cases | 2666 |
| Afghanistan | 2000 | population | 20595360 |
| Brazil | 1999 | cases | 37737 |
| Brazil | 1999 | population | 172006362 |
| Brazil | 2000 | cases | 80488 |
| Brazil | 2000 | population | 174504898 |
| China | 1999 | cases | 212258 |
| China | 1999 | population | 1272915272 |
| China | 2000 | cases | 213766 |
| China | 2000 | population | 1280428583 |
Tuberculosis data¶
This data is tidy. True or false?
| country | year | cases | population |
|---|---|---|---|
| Afghanistan | 1999 | 745 | 19987071 |
| Afghanistan | 2000 | 2666 | 20595360 |
| Brazil | 1999 | 37737 | 172006362 |
| Brazil | 2000 | 80488 | 174504898 |
| China | 1999 | 212258 | 1272915272 |
| China | 2000 | 213766 | 1280428583 |
Tools for tidying and wrangling data¶
tidyversepackage functions from:dplyrpackage (select,filter,mutate,group_by,summarize)tidyrpackage (pivot_longer,pivot_wider)purrrpackage (map_dfr)
Demo Time!¶

library(tidyverse)
library(palmerpenguins)
options(repr.matrix.max.rows = 6)
penguins
Select¶
The select function is used to select a subset of columns (variables) from a dataframe.
# select penguins example
Filter¶
The filter function is used to choose a subset of rows (observations) in a dataframe.
e.g. filter for only penguins with flippers longer than 190 mm
# penguins filter example
Mutate¶
The mutate function transforms old columns to add new columns.
e.g. convert body mass from grams to pounds
# penguins mutate example
Many operations in the same sequence¶
When you need type out a long sequence of operations on data, you could either:
- Save intermediate objects
- Compose function
- Use the pipe operator
|>
Let's look closer at each one of these
1. Save intermediate objects¶
penguins_1 <- mutate(penguins, body_mass_lb = body_mass_g/454)
penguins_2 <- filter(penguins_1, flipper_length_mm > 190)
penguins_3 <- select(penguins_2, bill_length_mm, bill_depth_mm, flipper_length_mm, body_mass_lb)
penguins_3
Disadvantages:¶
- The reader may be tricked into thinking the named
penguins_1andpenguins_2objects are important for some reason, while they are just temporary intermediate computations. - Further, the reader has to look through and find where
penguins_1andpenguins_2are used in each subsequent line. - Creating variables that we don't need, could lead to memory issues if they are big and take up a lot of space.
2. Composing functions¶
penguins_composed <- select(
filter(
mutate(
penguins, body_mass_lb = body_mass_g/454
),
flipper_length_mm > 190
),
bill_length_mm, bill_depth_mm, flipper_length_mm, body_mass_lb
)
penguins_composed
Disadvantage:¶
- Difficult to read since you the innermost function is exectued first
- You have to start reading from the middle of the code block (
mutateabove)
- You have to start reading from the middle of the code block (
3. Piping¶
You can also pipe with the |> symbol: passes the output of a function to the 1st argument of another.
penguins_piped <- penguins |>
mutate(body_mass_lb = body_mass_g/454) |>
filter(flipper_length_mm > 190) |>
select(bill_length_mm, bill_depth_mm, flipper_length_mm, body_mass_lb)
penguins_piped
note: if R sees a |> at the end of a line, it keeps reading the next line of code before evaluating!
Advantage:¶
- Pipes make code much more readable when you need to do a long sequence of operations on data.
- No intermediatery variables created
Nursery rhyme example¶
Jack and Jill went up the hill
To fetch a pail of water
Jack fell down and broke his crown
And Jill came tumbling after
Intermediate objects¶
on_hill <- went_up(jack_jill, 'hill')
with_water <- fetch(on_hill, 'water')
fallen <- fell_down(with_water, 'jack')
broken <- broke(fallen, 'jack')
after <- tumble_after(broken, 'jill')
Nursery rhyme example¶
Jack and Jill went up the hill
To fetch a pail of water
Jack fell down and broke his crown
And Jill came tumbling after
Pipes¶
jack_and_jill |>
went_up("hill") |>
fetch("water") |>
fell_down("jack") |>
broke("crown") |>
tumble_after("jill")
Credit: https://trestletech.com/wp-content/uploads/2015/07/dplyr.pdf
Grouping and summarizing via group_by + summarize¶
- Grouping is when you split data into groups based on the value of a column.
- Sumarizing is when you combine data into fewer summary values.
For example computing the average particle pollution per city (source https://info201.github.io/dplyr.html):
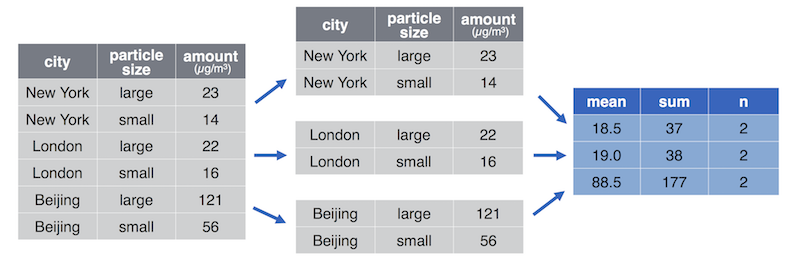
- Another example, splitting the
penguinsdata into one group perspecies, and then summarizing the values within each group e.g. reporting the averagebody_massper species.- To do this we use
group_by+summarizeto iterate over species, calculating average body mass.
- To do this we use
# Recall the data structure
penguins
# Calculate the average body mass for each species
What's going on with those NA's?
Another big concept this week: iteration¶
- Iteration is when you need to do something repeatedly (e.g., ringing in and bagging groceries at the till)

map_dfr¶
map_dfr iterates over columns,
e.g. to calculate the average for each numeric column in the penguins data
Be careful: There are a lot of map_... functions that you could use (map, map_lgl, map_chr, etc). Usually in this course we'll only use map_dfr, but you should investigate the others yourself!
# Calculate average each numeric column in penguins data set
Go forth and wrangle!¶

What did we learn?¶
Friends with similar tools:

Easier for automation & iteration!

And it makes all other tidy datasets seem more welcoming!

So make friends with tidy data!