set.seed(200)
sim_stats <- replicate(10000, {
x <- rnorm(100)
c(mean = mean(x),
median = median(x))
})
sim_stats <- t(sim_stats) |> as.data.frame()
sd(sim_stats$mean)[1] 0.1006556[1] 0.1[1] 0.1255247[1] 0.1253314DSCI 200
Katie Burak, Gabriela V. Cohen Freue
Last modified – 09 March 2026
\[ \DeclareMathOperator*{\argmin}{argmin} \DeclareMathOperator*{\argmax}{argmax} \DeclareMathOperator*{\minimize}{minimize} \DeclareMathOperator*{\maximize}{maximize} \DeclareMathOperator*{\find}{find} \DeclareMathOperator{\st}{subject\,\,to} \newcommand{\E}{E} \newcommand{\Expect}[1]{\E\left[ #1 \right]} \newcommand{\Var}[1]{\mathrm{Var}\left[ #1 \right]} \newcommand{\Cov}[2]{\mathrm{Cov}\left[#1,\ #2\right]} \newcommand{\given}{\ \vert\ } \newcommand{\X}{\mathbf{X}} \newcommand{\x}{\mathbf{x}} \newcommand{\y}{\mathbf{y}} \newcommand{\P}{\mathcal{P}} \newcommand{\R}{\mathbb{R}} \newcommand{\norm}[1]{\left\lVert #1 \right\rVert} \newcommand{\snorm}[1]{\lVert #1 \rVert} \newcommand{\tr}[1]{\mbox{tr}(#1)} \newcommand{\brt}{\widehat{\beta}^R_{s}} \newcommand{\brl}{\widehat{\beta}^R_{\lambda}} \newcommand{\bls}{\widehat{\beta}_{ols}} \newcommand{\blt}{\widehat{\beta}^L_{s}} \newcommand{\bll}{\widehat{\beta}^L_{\lambda}} \newcommand{\U}{\mathbf{U}} \newcommand{\D}{\mathbf{D}} \newcommand{\V}{\mathbf{V}} \]
Some examples and references are from Challenges of cellwise outliers, by Raymaekers and Rousseeuw
Robust Statistics, by Maronna, Martin, Yohai, Salibian-Barrera
In memory of my mentor and friend
Professor Ruben Zamar (1949-2023)
who contributed greatly to Robust Statistics and beyond …
By the end of this lesson, you will be able to:
Definition: Outlier
An outlier is an observation that deviates from the bulk of the data, an atypical observation.
errors when collecting or processing data
values generated from a different distribution
rare cases which may carry valuable information
As with missing values, outliers require careful diagnosis and appropriate handling before analysis.
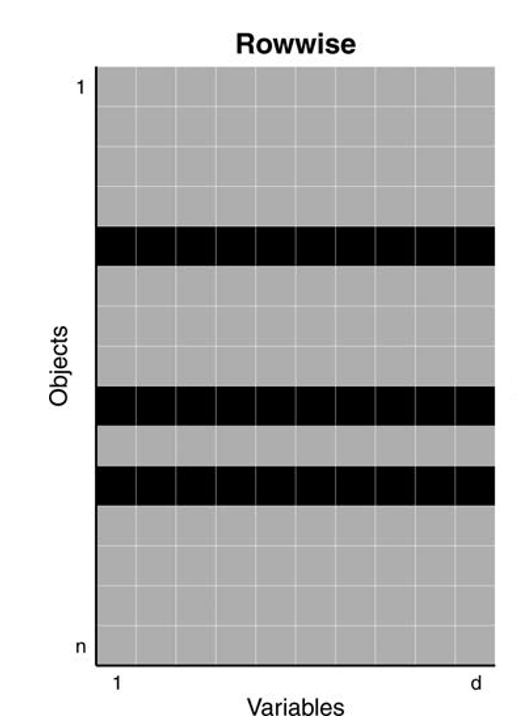
In the 1960s Tukey and Huber introduced the casewise (aka rowwise) contamination model
Treats entire rows as contaminated and coming from a different distribution, even if only values of some variables are unusual
Assumes that less than 50% of the cases (objects) are contaminated
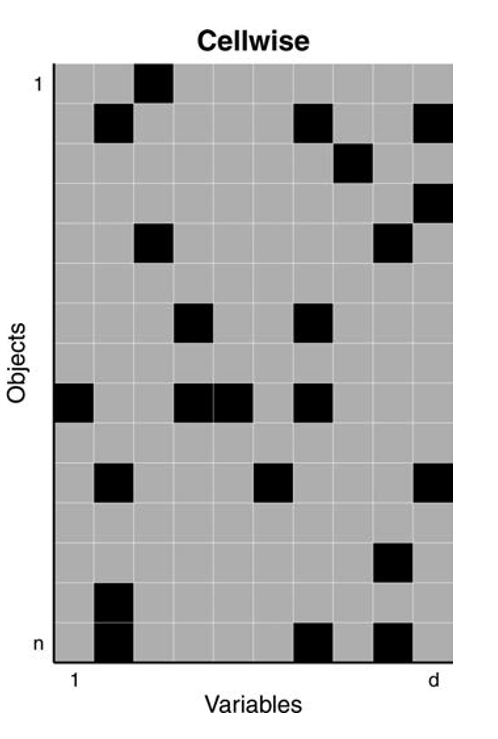
In 2009, Alqallaf, Van Aelst, Yohai and Zamar introduced the cellwise contamination model
Only individual cells are contaminated, with values coming from a different distribution
Any case (row) may contain some contaminated cells
More realistic for high-dimensional data
Data: n = 180 archeological glass spectra with d = 750 wavelengths (Detecting Deviating Data Cells, Rousseeuw & Van Den Bossche, Technometrics 2018)
Before analyzing multivariate datasets, we first need to learn how to identify and handle outlying values in a single variable.
Let’s look at the values of the wavelength V169 in this data
Classical estimators can be highly distorted by outliers
Robust estimators are needed to capture the bulk of the data
Robust estimators are also needed to flag outliers
| Statistic | Estimate |
|---|---|
| Mean | 5260.72 |
| Median | 4629 |
| SD | 1413.69 |
| MAD | 470.73 |
If the median is less affected by extreme values, why not using it as a default?
if the data contain outliers, the median is more resistant
but if the data does not contain outliers, it is less efficient (larger standard error!)
\[SE(\bar{X}) = \frac{\sigma}{\sqrt{n}}\]
\[SE(\tilde{X})= \sqrt{\frac{\pi}{2}}\frac{\sigma}{\sqrt{n}}\]
Let’s examine these two points by simulation!
Generate a sample of size 100 from a Normal distribution with mean 0 and standard deviation 1
Compute the mean and the median
Repeat 10000 times
Summarize their sampling distributions
Generate a sample of size 100, with 90% from \(\mathcal{N}(0,1)\) and 10% from \(\mathcal{N}(0,10)\)
Compute the mean and the median
Repeat 10000 times
Summarize their sampling distributions
\[\text{Var}(\bar{X}) = \frac{(1-\varepsilon)\sigma^2_1 + \varepsilon \sigma_2^2}{n}\]
\[\mathrm{Var}(\tilde{X}) \approx \frac{\pi}{2n}\; \frac{1}{\left(\frac{1-\varepsilon}{\sigma_1}+\frac{\varepsilon}{\sigma_2}\right)^2}\]
rcont_norm <- function(n, eps = 0.10, mu0 = 0,
sigma0 = 1, sigma1 = 10) {
x <- rnorm(n, mean = mu0, sd = sigma0)
k <- ceiling(eps * n)
if (k > 0) {
idx <- sample(1:n, k)
x[idx] <- rnorm(k, mean = mu0, sd = sigma1)
}
x
}
set.seed(200)
sim_stats <- replicate(10000, {
x <- rcont_norm(100)
c(mean = mean(x),
median = median(x))
})
sim_stats <- t(sim_stats) |> as.data.frame()
sd(sim_stats$mean)[1] 0.3307419[1] 0.1374634[1] 0.1377268Outliers can occur in all variables of an observation or only in certain variables
Cellwise paradigm is better is more flexible for high-dimensional data
Classical estimators can be optimal but very sensitive to outliers
Median is less efficient (≈ 1.57× variance) but more stable under contamination
Robustness = trade-off with efficiency
UBC DSCI 200